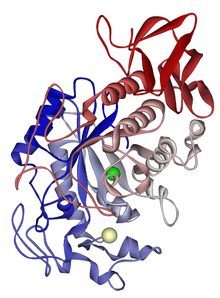
Alpha-amylase
| Alpha-amylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC number | 3.2.1.1 | ||||||||
| CAS number | 9000-90-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
| Alpha-amylase catalytic domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Cyclodextrin glucanotransferase (e.c.2.4.1.19) (cgtase) | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amylase | ||||||||
| Pfam | PF00128 | ||||||||
| Pfam clan | CL0058 | ||||||||
| InterPro | IPR006047 | ||||||||
| SCOP | 1ppi | ||||||||
| SUPERFAMILY | 1ppi | ||||||||
| OPM superfamily | 125 | ||||||||
| OPM protein | 1wza | ||||||||
| CAZy | GH13 | ||||||||
| |||||||||
| Alpha-amylase C-terminal beta-sheet domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Crystal structure of barley alpha-amylase isozyme 1 (amy1) inactive mutant d180a in complex with maltoheptaose | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amyl_C2 | ||||||||
| Pfam | PF07821 | ||||||||
| InterPro | IPR012850 | ||||||||
| |||||||||
| Alpha amylase, C-terminal all-beta domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 maltotriose complex of preconditioned cyclodextrin glycosyltransferase mutant | |||||||||
| Identifiers | |||||||||
| Symbol | Alpha-amylase_C | ||||||||
| Pfam | PF02806 | ||||||||
| Pfam clan | CL0369 | ||||||||
| InterPro | IPR006048 | ||||||||
| SCOP | 1ppi | ||||||||
| SUPERFAMILY | 1ppi | ||||||||
| |||||||||
α-Amylase is a protein enzyme EC 3.2.1.1 that hydrolyses alpha bonds of large, alpha-linked polysaccharides, such as starch and glycogen, yielding glucose and maltose.[2] It is the major form of amylase found in humans and other mammals.[3] It is also present in seeds containing starch as a food reserve, and is secreted by many fungi.
In human physiology
Although found in many tissues, amylase is most prominent in pancreatic juice and saliva, each of which has its own isoform of human α-amylase. They behave differently on isoelectric focusing, and can also be separated in testing by using specific monoclonal antibodies. In humans, all amylase isoforms link to chromosome 1p21 (see AMY1A).
Salivary amylase (ptyalin)
Amylase is found in saliva and breaks starch into maltose and dextrin. This form of amylase is also called "ptyalin" /ˈtaɪəlɪn/[4] It will break large, insoluble starch molecules into soluble starches (amylodextrin, erythrodextrin, and achrodextrin) producing successively smaller starches and ultimately maltose. Ptyalin acts on linear α(1,4) glycosidic linkages, but compound hydrolysis requires an enzyme that acts on branched products. Salivary amylase is inactivated in the stomach by gastric acid. In gastric juice adjusted to pH 3.3, ptyalin was totally inactivated in 20 minutes at 37 °C. In contrast, 50% of amylase activity remained after 150 minutes of exposure to gastric juice at pH 4.3.[5] Both starch, the substrate for ptyalin, and the product (short chains of glucose) are able to partially protect it against inactivation by gastric acid. Ptyalin added to buffer at pH 3.0 underwent complete inactivation in 120 minutes; however, addition of starch at a 0.1% level resulted in 10% of the activity remaining, and similar addition of starch to a 1.0% level resulted in about 40% of the activity remaining at 120 minutes.[6]
Optimum conditions for ptyalin
- Optimum pH - 7.0 [7]
- Human body temperature
- Presence of certain anions and activators:
Genetic variation in human salivary amylase
The salivary amylase gene has undergone duplication during evolution, and DNA hybridization studies indicate many individuals have multiple tandem repeats of the gene. The number of gene copies correlates with the levels of salivary amylase, as measured by protein blot assays using antibodies to human amylase. Gene copy number is associated with apparent evolutionary exposure to high-starch diets.[8] For example, a Japanese individual had 14 copies of the amylase gene (one allele with 10 copies, and a second allele with four copies). The Japanese diet has traditionally contained large amounts of rice starch. In contrast, a Biaka individual carried six copies (three copies on each allele). The Biaka are rainforest hunter-gatherers who have traditionally consumed a low-starch diet. Perry and colleagues speculated the increased copy number of the salivary amylase gene may have enhanced survival coincident to a shift to a starchy diet during human evolution.
Pancreatic amylase
Pancreatic α-amylase randomly cleaves the α(1-4) glycosidic linkages of amylose to yield dextrin, maltose, or maltotriose. It adopts a double displacement mechanism with retention of anomeric configuration.
In pathology
The test for amylase is easier to perform than that for lipase, making it the primary test used to detect and monitor pancreatitis. Medical laboratories will usually measure either pancreatic amylase or total amylase. If only pancreatic amylase is measured, an increase will not be noted with mumps or other salivary gland trauma.
However, because of the small amount present, timing is critical when sampling blood for this measurement. Blood should be taken soon after a bout of pancreatitis pain, otherwise it is excreted rapidly by the kidneys.
Salivary α-amylase has been used as a biomarker for stress that does not require a blood draw.[9]
Interpretation
Increased plasma levels in humans are found in:
- Salivary trauma (including anaesthetic intubation)
- Mumps — due to inflammation of the salivary glands
- Pancreatitis — because of damage to the cells that produce amylase
- Renal failure — due to reduced excretion
Total amylase readings of over 10 times the upper limit of normal (ULN) are suggestive of pancreatitis. Five to 10 times the ULN may indicate ileus or duodenal disease or renal failure, and lower elevations are commonly found in salivary gland disease.
Genes
In grain
α-Amylase activity in grain is measured by, for instance, the Hagberg-Perten Falling Number, a test to assess sprout damages,[10] or the Phadebas method.
Industrial use
α-Amylase is used in ethanol production to break starches in grains into fermentable sugars.
The first step in the production of high-fructose corn syrup is the treatment of cornstarch with α-amylase, producing shorter chains of sugars oligosaccharides.
An α-amylase called "Termamyl", sourced from Bacillus licheniformis, is also used in some detergents, especially dishwashing and starch-removing detergents.[11]
See amylase for more uses of the amylase family in general.
Buffer inhibition
The tris molecule is reported to inhibit a number of bacterial α-amylases,[12][13] so they should not be used in tris buffer.
Determination
Several methods are available for determination of α-amylase activity, and different industries tend to rely on different methods. The starch iodine test, a development of the iodine test, is based on colour change, as α-amylase degrades starch and is commonly used in many applications. A similar but industrially produced test is the Phadebas amylase test, which is used as a qualitative and quantitative test within many industries, such as detergents, various flour, grain, and malt foods, and forensic biology.
Domain architecture
α-Amylases contain a number of distinct protein domains. The catalytic domain has a structure consisting of an eight-stranded alpha/beta barrel that contains the active site, interrupted by a ~70-amino acid calcium-binding domain protruding between beta strand 3 and alpha helix 3, and a carboxyl-terminal Greek key beta-barrel domain.[14] Several alpha-amylases contain a beta-sheet domain, usually at the C terminus. This domain is organised as a five-stranded antiparallel beta-sheet.[15][16] Several alpha-amylases contain an all-beta domain, usually at the C terminus.[17]
See also
References
- ↑ Ramasubbu, N.; Paloth, V.; Luo, Y.; Brayer, G. D.; Levine, M. J. (1996). "Structure of Human Salivary α-Amylase at 1.6 Å Resolution: Implications for its Role in the Oral Cavity". Acta Crystallographica Section D. 52 (3): 435–446. doi:10.1107/S0907444995014119. PMID 15299664.
- ↑ Maureen Barlow Pugh, ed. (2000). Stedman's Medical Dictionary (27th ed.). Baltimore, Maryland, USA: Lippincott Williams & Wilkins. p. 65. ISBN 978-0-683-40007-6.
- ↑ Voet, D., & Voet, J. G. (2005). Biochimie. (2e éd.). Bruxelles: De Boeck. 1583 p.
- ↑ which was named by Swedish chemist Jöns Jacob Berzelius. The name derives from the Greek word πτυω (I spit), because the substance was obtained from saliva. See: J. Berzelius (Ms. Esslinger, trans.), Traité de Chimie (Paris, France: Firmin Didot Frerès, 1833), vol. 7, page 156.
- ↑ Fried, M, Abramson, S, Meyer, JH. Passage of salivary amylase through the stomach in humans. Digestive Diseases and Sciences 32:1097-1103 (1987).
- ↑ Rosenblum, JL, Irwin, CI, Alpers, DH. Starch and glucose oligosaccharides protect salivary-type amylase activity at acid pH. American Journal of Physiology 254 (Gastrointestinal and Liver Physiology 17):G775-780 (1988).
- ↑ http://www.worthington-biochem.com/aa/
- ↑ Perry, GH, et al. Diet and evolution of human amylase gene copy number variation, Nature Genetics 39:1256-1260 (2007).
- ↑ Noto, Yuka; Tetsumi Sato; Mihoko Kudo; Kiyoshi Kurata; Kazuyoshi Hirota (December 2005). "The Relationship Between Salivary Biomarkers and State-Trait Anxiety Inventory Score Under Mental Arithmetic Stress: A Pilot Study". Anesthesia & Analgesia. United States: Lippincott Williams & Wilkins. 101 (6): 1873–6. doi:10.1213/01.ANE.0000184196.60838.8D. PMID 16301277. Retrieved 1 November 2013.
- ↑ "Falling Number – Introduction". Perten Instruments. 2005. Retrieved 21 November 2009.
- ↑ "The use of enzymes in detergents". Faculty of Engineering, Science and the Built Environment, London South Bank University. 20 December 2004. Archived from the original on 20 October 2009. Retrieved 21 November 2009.
- ↑ Ghalanbor, Z; et al. (2008). "Binding of Tris to Bacillus licheniformis alpha-amylase can affect its starch hydrolysis activity.". Protein Peptide Lett. 15 (2): 212–214. doi:10.2174/092986608783489616. PMID 18289113.
- ↑ Aghajari, N; et al. (1998). "Crystal structures of the psychrophilic alpha-amylase from Alteromonas haloplanctis in its native form and complexed with an inhibitor.". Protein Sci. 7 (3): 564–572. doi:10.1002/pro.5560070304. PMC 2143949
 . PMID 9541387.
. PMID 9541387. - ↑ Abe A, Yoshida H, Tonozuka T, Sakano Y, Kamitori S (December 2005). "Complexes of Thermoactinomyces vulgaris R-47 alpha-amylase 1 and pullulan model oligossacharides provide new insight into the mechanism for recognizing substrates with alpha-(1,6) glycosidic linkages". FEBS J. 272 (23): 6145–53. doi:10.1111/j.1742-4658.2005.05013.x. PMID 16302977.
- ↑ Kadziola A, Søgaard M, Svensson B, Haser R (April 1998). "Molecular structure of a barley alpha-amylase-inhibitor complex: implications for starch binding and catalysis". J. Mol. Biol. 278 (1): 205–17. doi:10.1006/jmbi.1998.1683. PMID 9571044.
- ↑ Kadziola A, Abe J, Svensson B, Haser R (May 1994). "Crystal and molecular structure of barley alpha-amylase". J. Mol. Biol. 239 (1): 104–21. doi:10.1006/jmbi.1994.1354. PMID 8196040.
- ↑ Machius M, Wiegand G, Huber R (March 1995). "Crystal structure of calcium-depleted Bacillus licheniformis alpha-amylase at 2.2 A resolution". J. Mol. Biol. 246 (4): 545–59. doi:10.1006/jmbi.1994.0106. PMID 7877175.
External links
- The alpha-Amylase Protein
- alpha-Amylase at the US National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the public domain Pfam and InterPro IPR006047 This article incorporates text from the public domain Pfam and InterPro IPR012850 This article incorporates text from the public domain Pfam and InterPro IPR006048